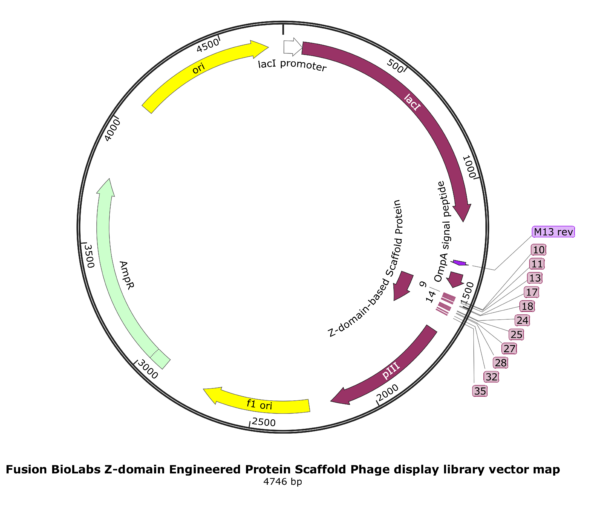
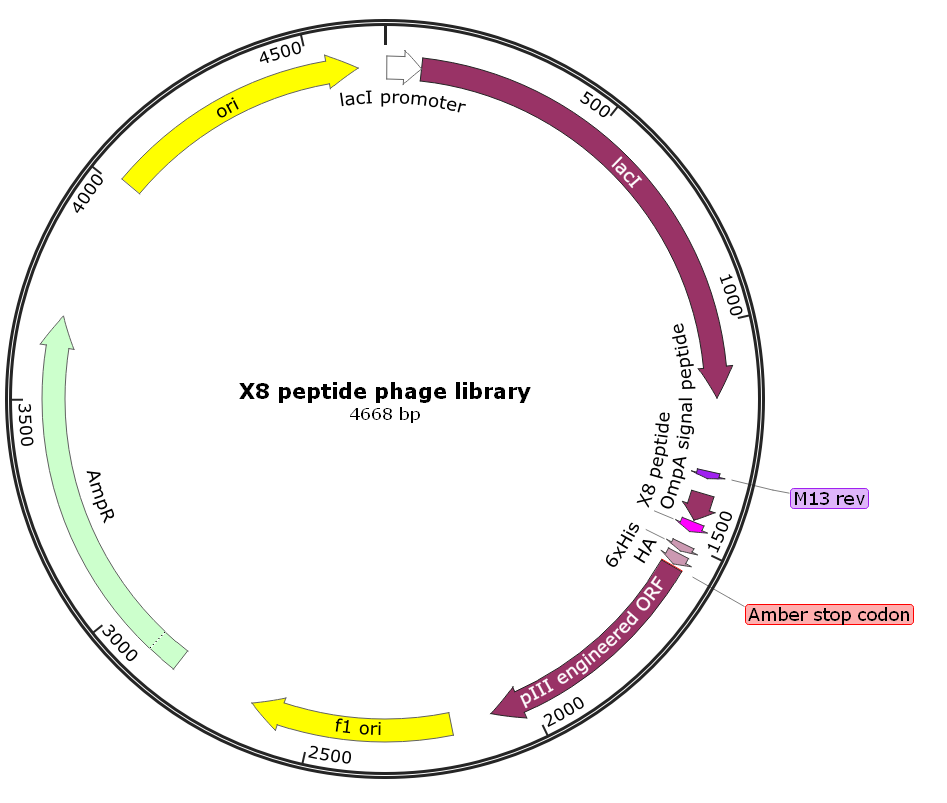
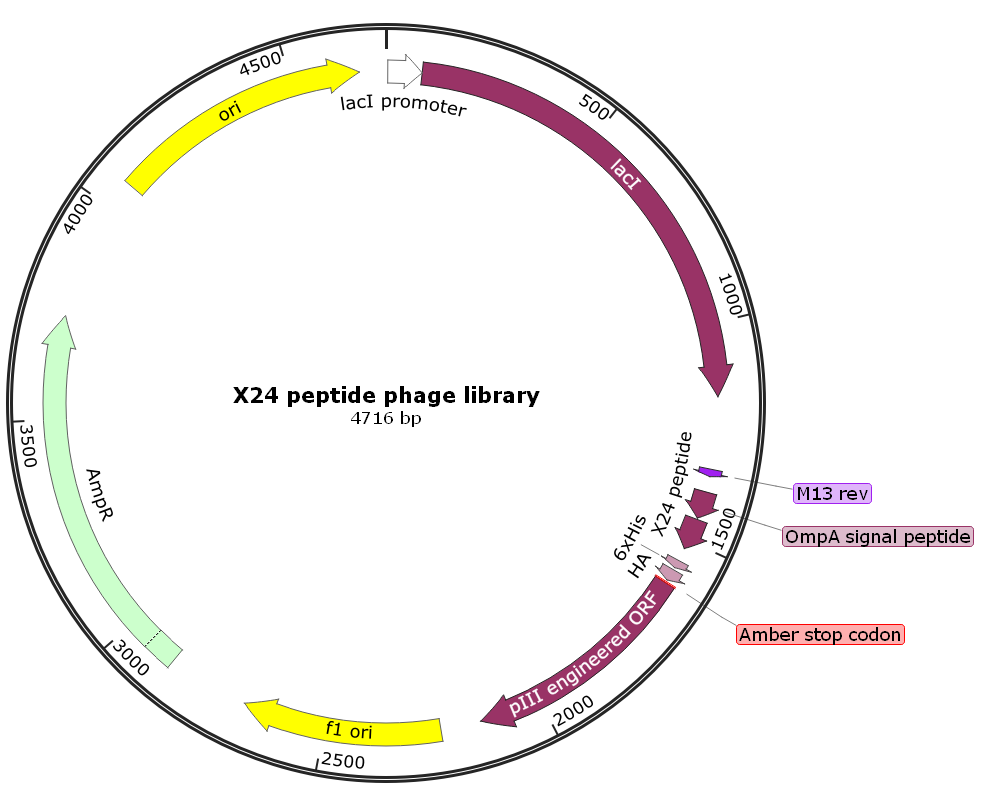
A Z-domain engineered scaffold protein is a small, highly stable, engineered protein derived from the “B domain” of staphylococcal protein A. It is a primary example of an “alternative scaffold”—a protein used as a framework to create new binding molecules that can perform similar functions to antibodies but with distinct physical advantages. This library can be screened to obtain Z-domain engineered protein scaffold molecules with high affinity and specificity to any given target molecules.
The Z-domain is a highly versatile engineered scaffold protein. It is primarily used as a robust structural framework for developing novel binding molecules.
- Origin: It is an engineered analog of the B-domain of Staphylococcal protein A.
- Structure: It consists of 58 amino acids organized into a stable three-helix bundle.
- Natural Affinity: Its parent form (Z-IgG) binds specifically to the Fc region of IgG antibodies.
- Engineered Versatility: By randomizing 13 surface-exposed residues on the first two helices, researchers can create variants that bind to diverse targets, such as HER2, TNFα, and VEGF.
- Stability: It is remarkably stable and can refold easily after denaturation.
- Simplicity: Its small size makes it easy to produce in E. coli and ideal for chemical synthesis.
- Durability: It is tolerant to many substitutions, allowing for a wide range of “patch engineering” to create new binding sites.
The Fusion BioLabs Z-domain engineered protein scaffold ready-to-panning phage display library was constructed by saturation mutagenesis of the Z domain at 13 solvent-exposed positions (9, 10, 11, 13, 14, 17, 18, 24, 25, 27, 28, 32 and 35) across Helices 1 and 2.
VDNKFNKEXXXAXXEIXXLPNLNXXQXXAFIXSLXDDPSQSANLLAEAKKLNDAQAPK
- Format: OmpA leader sequence-Z-domain engineered protein scaffold
- Diversity: 4.8×1011
- The diversity and in-frame of the library were checked by NGS
Z-domain engineered protein scaffold Library
| Library Description | Format | Diversity/Phage titer (pfu/ml) checked by NGS | |
|---|---|---|---|
| Z-domain Engineered Protein Scaffold | aturation mutagenesis at 13 solvent-exposed positions (9, 10, 11, 13, 14, 17, 18, 24, 25, 27, 28, 32, and 35) across Helices 1 and 2 | OmpA leader sequence—Z-domain engineered protein scaffold | 4.8×1011/9.2×1012 |
- Kit contents
The following components are included in the kit.
Ready-to-panning Z-domain-based Scaffold Protein Library Kit Content
| Component | Quantity | Composition |
|---|---|---|
| Z-domain-based scaffold protein library (ready-to-panning phage; 9.2×1012pfu/ml) | 1.0 ml | 1×PBS with 25% glycerol |
| M13KO7 Helper Phage (2.0×1012 pfu/ml) | 0.5 ml | 1×PBS with 25% glycerol |
| Chemically Competent TG1 E. coli (2.5x108cfu/ml) | 0.5 ml | 2xYT with 25% Glycerol |
| M13 Reverse Primer (1.6 µM) | 0.2 ml | 1×TE Buffer |
Key Advantages
- Small Size (~6.5 kDa): Ideal for tissue penetration and applications where large antibodies fail.
- Robust Folding: Rapid, reversible folding without the need for disulfide bonds; compatible with reducing environments.
- Exceptional Stability: Resistant to high temperatures and a wide range of pH levels (pH 2–11).
- Clean Intellectual Property: Based on established, public-domain Z-domain sequences. Avoids the royalties and restrictions associated with proprietary “next-generation” stabilized scaffolds.
Applications
- Alternative Scaffold Discovery: Identify novel binding proteins for diagnostic imaging agents.
- Research Tools: Create high-affinity ligands for protein purification and Western blotting.
- Biophysical Studies: Study protein-protein interactions using a stable, minimal framework.
Frequently Asked Questions (FAQ)
- Is this an Affibody® library?
No. This is a Z-domain-based scaffold protein library. While it shares a similar structural lineage (the 3-helix bundle derived from Protein A), it uses the original public-domain Z-domain sequence and the classic 13-position randomization map. We are not affiliated with Affibody AB, and our products are not licensed Affibody® molecules.
- What is the difference between this and “optimized” scaffolds?
Proprietary “optimized” scaffolds (such as those used in clinical-stage candidates like izokibep) often contain additional mutations (e.g., at positions 23, 33, or 43) to enhance thermal stability or reduce antibody binding. Our library uses the high-performance original Z-domain backbone (starting with VDNK…). This version is robust, highly stable, and operates under a clear, non-proprietary IP framework.
- Can I use the binders I find for commercial purposes?
Our library is sold for Research Use Only (RUO). While the scaffold itself is based on public-domain sequences, the specific binding sequences you discover against certain medical targets (like HER2, IL-17, or TNFα) may be subject to third-party patents held by other companies. We recommend performing a Freedom to Operate (FTO) search before moving a specific lead into clinical or commercial development.
- Why should I use a Z-domain library instead of an antibody library?
Z-domain scaffolds offer several advantages over traditional IgG libraries:
- Size: At 6.5 kDa, they are 25x smaller than antibodies, allowing for superior tissue penetration.
- Stability: They fold correctly in the cytoplasm (no disulfide bonds needed) and withstand temperatures up to 75°C.
- Production: They are easily expressed at high levels in E. coli, significantly reducing manufacturing costs.
Comparison: Z-Domain Scaffolds vs. Conventional Antibodies (mAbs)
| Feature | Z-Series™ Z-Domain Library | Conventional Antibodies (IgG) |
|---|---|---|
| Molecular Weight | ~6.5 kDa (Ultra-small) | ~150 kDa (Large) |
| Structure | 3-Helix Bundle (58 amino acids) | 4-Chain Complex (Y-shape) |
| Disulfide Bonds | None (Folds in reducing environments) | Required (Complex folding) |
| Tissue Penetration | Excellent (Rapid & Deep) | Poor (Slow diffusion) |
| Thermal Stability | High (Resistant up to 75°C) | Moderate (Sensitive to heat) |
| pH Stability | Broad (pH 2 – 11) | Narrow (Sensitive to acid/base) |
| Production Host | E. coli (Simple & Inexpensive) | Mammalian Cells (Complex & Costly) |
| Blood Clearance | Rapid (Ideal for high-contrast imaging) | Slow (Days to weeks) |
| Immunogenicity | Generally Low | Variable (Depends on "humanization") |
| Engineering Ease | Simple (Single-gene fusion) | Complex (Heavy & Light chains) |
- Why Choose the Z-Domain Library?
- Deep Tissue Imaging: Due to their size (1/25th of an antibody), Z-domain binders can penetrate tumors or dense tissues much faster, providing clearer diagnostic results within hours rather than days.
- Intracellular Applications: Because they do not require disulfide bonds, these proteins fold correctly inside the cytoplasm of living cells—a feat most antibodies cannot achieve.
- Cost-Efficient Scale-Up: Since they are produced in E. coli, the cost per milligram is significantly lower than monoclonal antibodies produced in CHO (Chinese Hamster Ovary) cells.
- Stable Under Stress: They remain functional after exposure to harsh conditions (like those used in chemical conjugation or radiolabeling), where traditional antibodies would denature.
- Is the library compatible with standard phage display protocols?
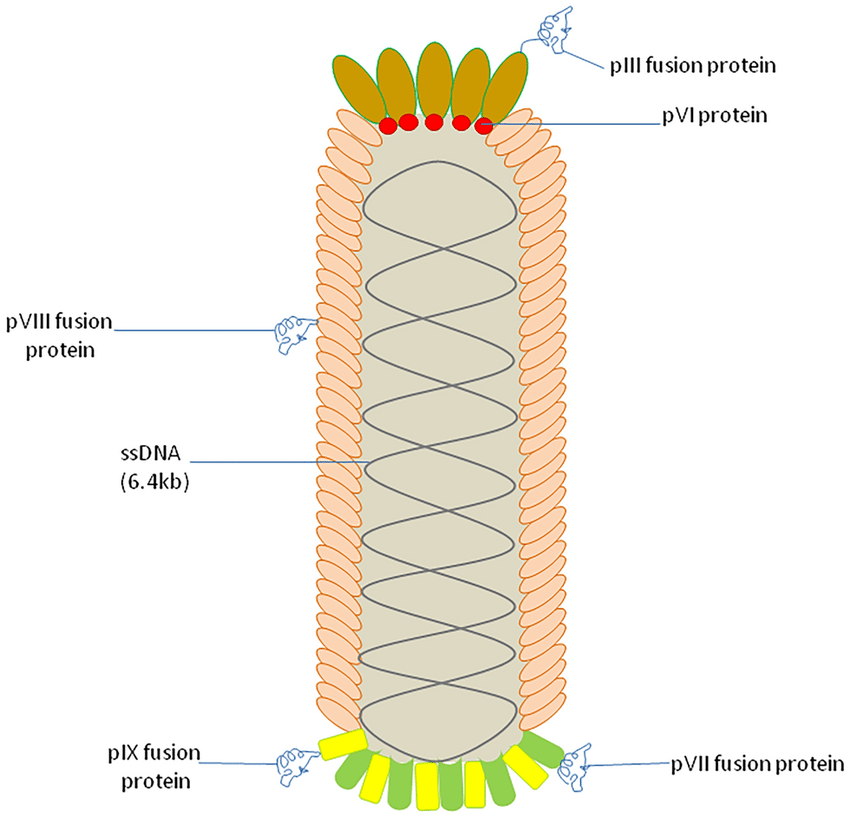
Yes. Our library is provided in a standard phagemid format (M13 pIII fusion). It is compatible with conventional panning techniques used for ScFv or VHH libraries. We provide a detailed technical protocol with every purchase to ensure a smooth workflow.
- Can these proteins be radiolabeled or conjugated?
Absolutely. The Z-domain scaffold is remarkably resilient to the harsh chemical conditions often required for radiolabeling (for PET/SPECT imaging) or site-specific conjugation to toxins and fluorophores.
Disclaimer for Terms of Sale
INTELLECTUAL PROPERTY AND USE LIMITATIONS
- Nature of Product: This product is a generic Z-domain-based combinatorial protein library derived from the B-domain of Staphylococcal protein A. This product is not an “Affibody®” molecule, and Fusion BioLabs is not affiliated with, endorsed by, or licensed by Affibody AB.
- No Trademark License: The purchase of this library does not grant the buyer any right or license to use the trademark “Affibody®” or any other protected trademarks in connection with the buyer’s research, products, or commercial activities.
- Freedom to Operate (FTO): While the scaffold backbone provided is based on public domain sequences, Fusion BioLabs makes no warranty that the use of this library to identify binders for specific biological targets (e.g., IL-17, HER2, TNFα, etc.) does not infringe upon third-party patents.
- Buyer Responsibility: The Buyer assumes sole responsibility for performing necessary patent searches and obtaining any required licenses for specific therapeutic or diagnostic targets discovered using this library.
- Research Use Only (RUO): This library and its derivatives are intended strictly for in vitro or in vivo laboratory research purposes. They are not approved for human diagnostic or therapeutic use.
- Limitation of Liability: In no event shall Fusion BioLabs be liable for any claims, losses, or damages resulting from the Buyer’s infringement of third-party intellectual property rights arising from the use of the library or the commercialization of molecules derived therefrom.
Reference:
Nord K, Gunneriusson E, Ringdahl J, Ståhl S, Uhlén M, Nygren PA. Binding proteins selected from combinatorial libraries of an alpha-helical bacterial receptor domain. Nat Biotechnol. 1997 Aug;15(8):772-777.